Jun 19, 2020 |
Introduction
Recognizing that it’s been a few years since many of us sat through a biology class, and given that we are all now thinking about the role(s) we might play in minimizing the impact of COVID-19, we thought a short primer on COVID-19 genetics might be of interest. By providing informaticians with a deeper understanding of viral biology, perhaps we can stimulate additional out-of-the box thinking.
First, a little nomenclature. The virus responsible for Coronavirus Disease 2019 (COVID-19) is SARS-CoV-2 (previously known as 2019-nCoV). SARS-CoV-2 is ‘phylogenetically similar’ to other coronaviruses SARS-CoV and MERS-CoV, both of which cause severe respiratory disease; and coronavirus OC43, HKU1, NL63, and 229E, which produce generally mild common cold symptoms.
Phylogenetic similarity is calculated based on a comparison of nucleotides or amino acids across different strains or specimens. It is the technique used, for instance, to show the similarity between SARS-CoV-2 and a coronavirus endemic to bats, leading to the conclusion that mutations in the bat coronavirus allowed it to make the leap to become infectious in humans. It is also a technique used to infer the temporal and geographic spread of the virus. SARS-CoV-2 viruses, like cancer cells, continue to mutate as they replicate, at a rate estimated to be two to four times slower than the flu. Genetic comparisons between cases can lead epidemiologists to conclude patterns of spread. Here for instance, is a phylogenetic tree of SARS-CoV-2 taken from nextstrain.org showing evolution of the virus over time, grouped by ‘clade’ (a clade is a grouping based on common ancestry). Of note is the common ancestry of early cases concentrated in the state of Washington, implying a common source, and a different ancestry of early cases in New York, implying a different source.
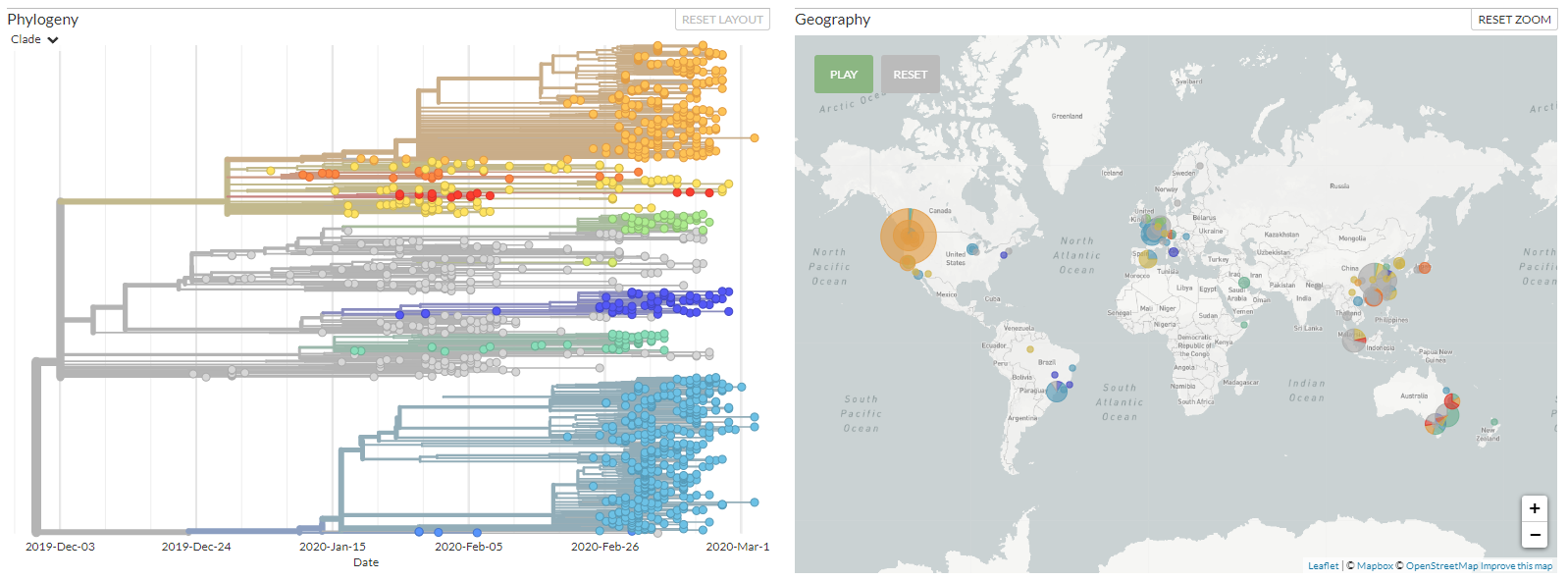
So, what exactly is a virus? And specifically, what are the genetics of SARS-CoV-2? Most simply, a virus is a small infectious agent that replicates only inside the living cells of a host organism. A complete virus particle, known as a virion, consists of a nucleic acid (DNA or RNA) surrounded by a protective coat of protein called a capsid.
A SARS-CoV-2 virion is a pleomorphic spherical particle, diameter around 120nm, with bulbous surface projections apparent under electron microscopy, reminiscent of the Latin corona, meaning ‘crown’ or ‘halo’, hence the virus’ name. They are enveloped viruses with a single-stranded ‘positive-sense RNA’ (i.e. RNA that can function directly as messenger RNA) genome, approximately 30 kilobases long. The viral envelope consists of a lipid bilayer where the membrane (M), envelope (E) and spike (S) structural proteins are anchored. Inside the envelope, there is the nucleocapsid, which is formed from multiple copies of the nucleocapsid (N) protein, and is bound to the RNA. (Figure is from here).
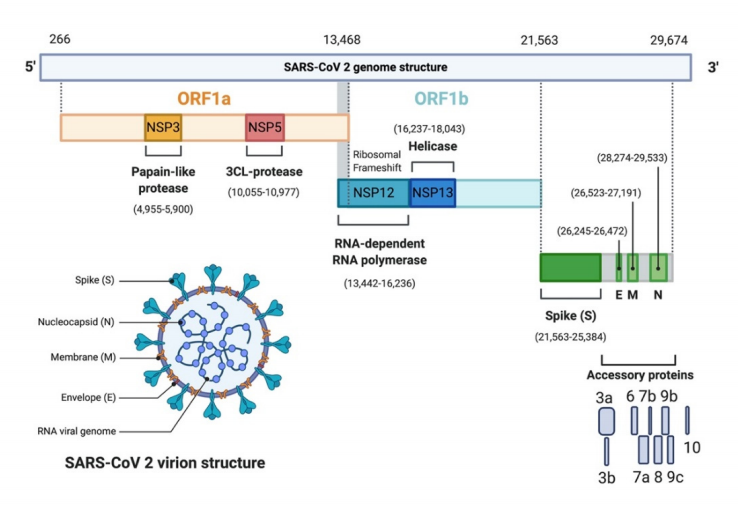
The life cycle of SARS-CoV-2 begins when the S protein (specifically, the Receptor Binding Domain of the S protein) attaches to a host cell’s ACE2 receptor, after which the viral envelope fuses with the cell membrane and releases the viral genome into the target cell.
For all you may have learned about viruses, SARS-CoV-2, and other positive-sense single-stranded RNA viruses are interesting in that they do not enter the cell’s nucleus. Rather, viral proteins and viral RNA are produced in the cell’s cytoplasm. During viral replication, the ORF1a and ORF1b genes of the viral RNA are first translated to produce many non-structural proteins that come together to produce a replicase-transcriptase enzyme complex that can both replicate (i.e. create an complementary copy of the viral RNA molecule) and transcribe (i.e. create shorter messenger RNA molecules that direct the translation of new structural proteins) the viral genome. New proteins assemble with new RNA molecules, and progeny viruses are then released from the host cell by exocytosis through secretory vesicles. The mechanism by which this life cycle triggers such aggressive inflammation and acute lung injury is under intense investigation.
Also under intense investigation is the spike (S) protein, which is crucial in determining cell tropism (which type of host tissue can be infected) and host species specificity (which host species can be infected). Mutations in the gene coding for the S protein may explain how the virus was able to become infective in humans, by enabling enhanced ACE2 receptor binding. In addition, preliminary evidence suggests that acquired mutations in the S protein affect viral pathogenicity. As we’ll discuss in greater detail below, human antibody production is often directed at the S protein, and may neutralize the virus.
Molecular Biology of Diagnosis and Treatment
Anything we write about diagnosis and treatment will become quickly out of date. Our intent here is to familiarize readers with how the viral biology is being exploited by informaticists seeking to discover new treatments. Readers are encouraged to consult more up to date sources for the most recent diagnostic and treatment information.
Diagnostic testing is primarily based on whole genome sequencing (WGS), real-time reverse transcriptase PCR (rRT-PCR), and antibody testing. WGS is more used for research and phylogenetic analysis, while rRT-PCR is used for detecting a current infection, and antibody testing is used for detecting evidence of a prior infection. rRT-PCR is a laboratory technique combining reverse transcription of RNA into DNA followed by amplification of specific DNA targets using polymerase chain reaction (PCR). It is primarily used to measure the amount of a specific RNA. Plus, with the addition of fluorescent technology, as the number of DNA amplification copies increases during the PCR reaction, there is also an increase in the fluorescence, offering both real time monitoring and quantitative analysis. Antibody testing detects a host’s antibody response to infection. IgG and IgM antibodies generally become detectable 1-3 weeks after symptom onset, at which time evidence suggests that infectiousness may be greatly decreased and that some degree of immunity from future infection has developed. The two major antigenic targets of SARS-CoV-2 virus against which antibodies are detected are the S and N proteins. In fact, the Receptor Binding Domain of the S protein (the portion of the S protein that interacts with the ACE2 receptor) is an immunodominant and highly specific target of antibodies. Evidence suggests that antibodies directed against this domain are neutralizing (i.e. they block the virus from entering human cells). While antibody testing is not currently being used to determine protective immunity and infectiousness among persons recently infected, not surprisingly, many experimental vaccine strategies center around stimulating the production of such antibodies.
Vaccine development is being aggressively pursued. Among the vaccine technologies under evaluation are whole virus vaccines, recombinant protein subunit vaccines, and nucleic acid vaccines. A major advantage of whole virus vaccines is their inherent immunogenicity. However, live virus vaccines often require extensive additional testing to confirm their safety. This is especially an issue for coronavirus vaccines, given the findings of increased infectivity following immunization with live or killed whole virus SARS coronavirus vaccines. Many subunit vaccines are directed at eliciting neutralizing antibodies to the SARS-CoV-2 S protein, in order to interfere with the ability of the virus to interact with the ACE2 receptor. Even more specifically, subunit vaccines that elicit antibodies focused against the Receptor Binding Domain of the S protein may elicit protective immunity while minimizing host immunopotentiation. Several promising trials are now showing production of neutralizing antibodies directed against the S protein.
There are currently no FDA-approved treatments for coronavirus.The drug farthest along in clinical trials is remdesivir. Most of the drugs in clinical trials inhibit key components of the coronavirus life cycle. These include viral entry into the host cell (blocked by umifenovir, chloroquine or interferon), viral replication (blocked by lopinavir/ritonavir, ASC09 or darunavir/cobicistat) and viral RNA synthesis (inhibited by remdesivir, favipiravir, emtricitabine/tenofovir alafenamide or ribavirin). Targeting viral cellular entry via the S protein, and targeting transmembrane serine protease TMPRSS2 activity are under investigation, as is the Janus-associated kinase (JAK) inhibitor Olumiant (baricitinib). Additional clinical trials are investigating drugs that alter the host’s immune response in an attempt to decrease the COVID-19 induced ‘cytokine storm‘.
Readers can get a sense of the unprecedented research in this area by visiting ClinicalTrials.gov, to see the multitude of active trials.
What can Informaticists do?
We were encouraged to write this piece by a number of our colleagues that felt that a primer on viral biology would stimulate further out-of-the-box thinking in the fight against COVID-19. And there is no doubt that knowledge is empowering – for instance, baricitinib was identified as a potential SARS-CoV-2 treatment using machine learning algorithms on the basis of its inhibition of ACE2-mediated endocytosis. That said, it’s also clear that ‘bread and butter’ informatics, such as described by Reeves, et al in their recent JAMIA piece, plays a crucial role, particularly when aligned with recommendations from established agencies such as the WHO and CDC. The EHR is a foundational public health tool, where informaticists can deploy clinical decision support, screening questionnaires, care plans and guidelines, and order sets; and where informaticists play a key role in standardized coding for aggregate knowledge management and real time data analytics. And the list goes on and on. It is likely that in the future, informaticists will discover viral variants that confer drug susceptibility or resistance, and human variants that influence antiviral therapy.
Conclusion
As we write this piece, the world has surpassed 8 million known cases with nearly half a million deaths attributable to COVID-19. We hope we’ve inspired some of you to take the time to learn more about the virus as one way of getting your own creative juices flowing. It’s likely that many of us can help save lives and lessen the global impact of the pandemic.
